Overview of research
Designing improved antifungal peptides
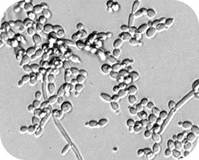
Antimicrobial peptides are short peptides that inhibit the growth of microorganisms, including bacteria and fungi. While the effect of antimicrobial peptides on cells is their most important characteristic, the stability of the peptides in their environment is also important to understand for developing peptides with therapeutic potential. We are studying the proteolytic stability of peptides with activity against the most common fungal pathogen, Candida albicans. One peptide we study is the human salivary antifungal peptide histatin 5. Histatin 5 has potent antifungal activity against C. albicans, but the fungus produces secreted aspartic proteases (Saps) that can degrade the peptide and make it inactive. We are designing modified peptides with reduced reduced cleavage by the Saps and improved antifungal activity. Engineering cell-penetrating peptides to target fungal pathogens Cell-penetrating peptides (CPPs) are short, structurally diverse peptides with the ability to cross cell membranes. While translocating into cells, they can carry a variety of bioactive cargo, including nucleic acids, proteins, and nanoparticles. CPPs have been widely studied in mammalian cells, but knowledge of their interactions with fungal cells are much more limited. We use biological experiments, biophysical experiments, and molecular simulations to study how the structure of CPPs affects their ability to translocate into Candida pathogens and their specificity for fungal cells versus mammalian cells. We are also studying how the properties of the molecular cargo carried by the CPPs affects translocation. Understanding protein-lipid interactions Lipid transport proteins are involved in the movement between lipid membranes in cells, and their dysfunction can lead to a number of different diseases. One important model lipid transport protein is the Osh4 protein in Saccharomyces cerevisiae, which is a homolog of a lipid transport protein in humans. Our lab is working with Prof. Jeff Klauda's lab to understand the molecular mechanism for how Osh4 interacts with membranes and the role it plays in moving lipids between membranes. We combine the molecular simulations performed by Prof. Klauda's lab with experiments performed in our lab to understand how lipid and protein properties affect the interaction of Osh4 with membranes. Engineering intracellular antibodies Intracellular antibodies ("intrabodies") are antibodies that bind to a target molecule inside cells. They must be engineered to fold in the reducing intracellular environment without the disulfide bonds normally required for proper folding. Although intrabodies have promise in therapeutics and research, the ability to efficiently engineer them to bind a target in the intracellular environment has limited their use. We developed and use an intrabody engineering method that takes advantage of an intracellular folding quality control mechanism in Escherichia coli to efficiently engineer intrabodies for proper intracellular folding and for binding to a target protein. We are using our screening method to engineer antibodies and other binding scaffolds with high affinity for targets that have biomedical relevance, including proteins important in cancer. |
|
|